Publications
Browse our recent published articles in peer-reviewed journals.2025

Spatial multi-omics reveals cell-type-specific nuclear compartments
Nature, 2025
Spatial multi-omics of nuclear architecture with two-layer seqFISH+
Yodai Takei
2024

Spatial Transcriptomics Reveals the Temporal Architecture of the Seminiferous Epithelial Cycle and Precise Sertoli-Germ Synchronization
BioRxiv, 2024
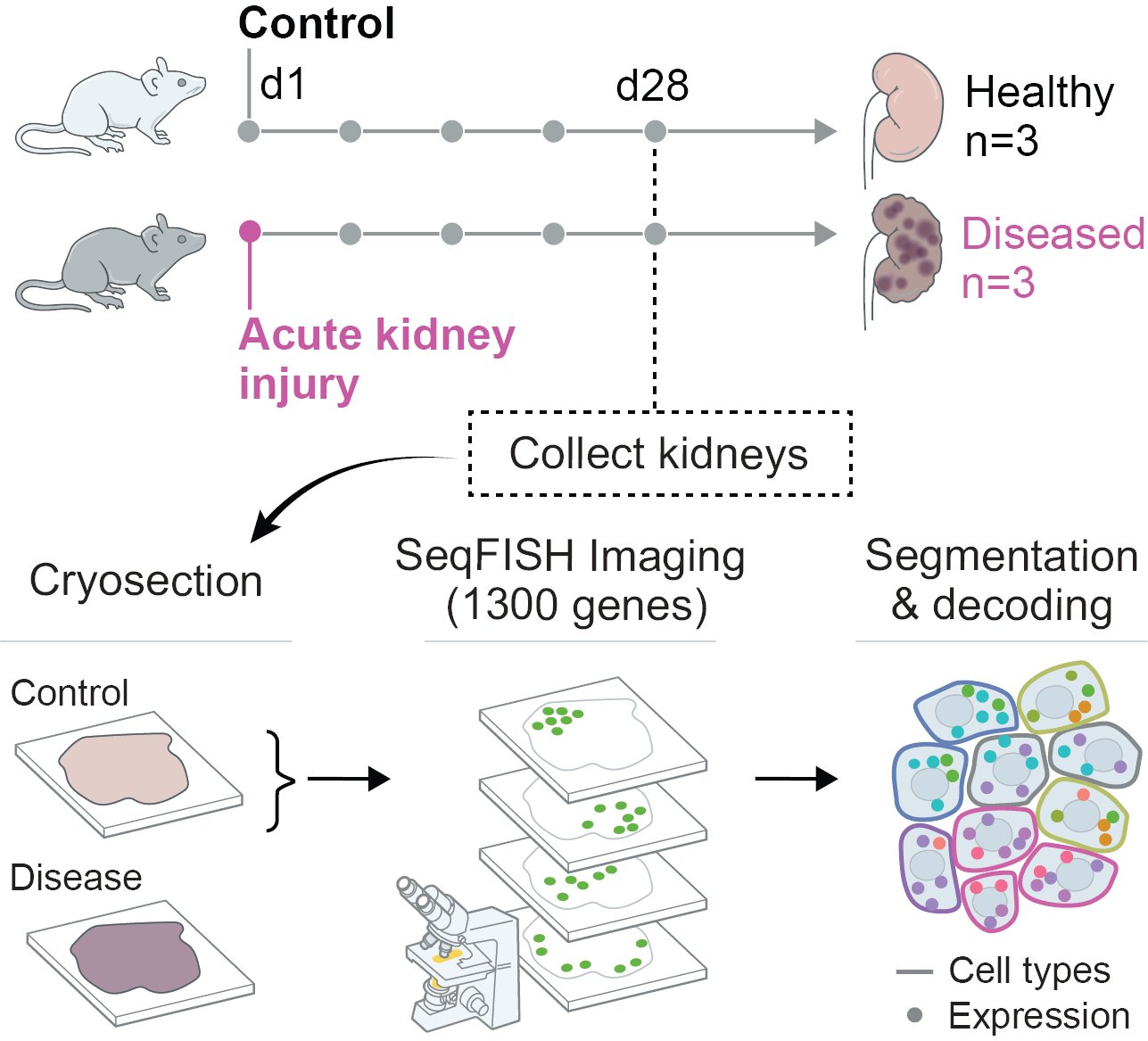
Spatial transcriptomics defines injury-specific microenvironments in the adult mouse kidney and novel cellular interactions in regeneration and disease
Nature Communications, 2024
2023

High-resolution spatial multi-omics reveals cell-type specific nuclear compartments
BioRxiv (2023)
2021
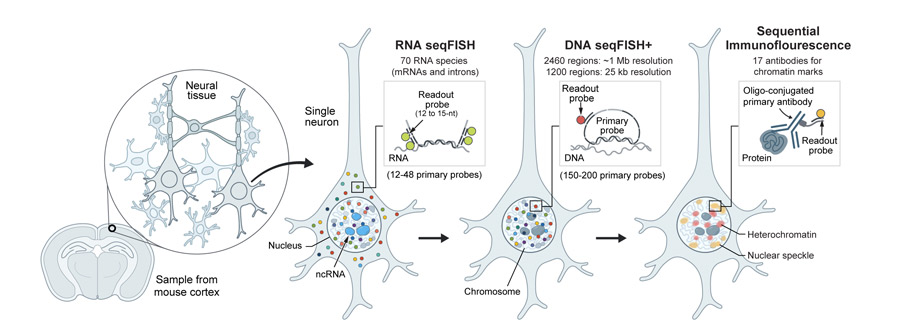
Single-cell nuclear architecture across cell types in the mouse brain
Science (2021)

Integrated spatial genomics reveals global architecture of single nuclei
Nature, (2021)
Yodai Takei, Jina Yun, Shiwei Zheng, Noah Ollikainen, Nico Pierson, Jonathan White, Sheel Shah, Julian Thomassie, Shengbao Suo, Chee-Huat Linus Eng, Mitchell Guttman, Guo-Cheng Yuan & Long Cai
Preprint (2020) at biorxiv.org
2020

Detecting protein and post-translational modifications in single cells with iDentification and qUantification sEparaTion (DUET)
Nature, Communications Biology (2020)
Yandong Zhang, Changho Sohn, Seoyeon Lee, Heejeong Ahn, Jinyoung Seo, Junyue Cao and Long Cai
2019

Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH+
Nature (2019)
Chee-Huat Linus Eng, Michael Lawson, Qian Zhu, Ruben Dries, Noushin Koulena, Yodai Takei, Jina Yun, Christopher Cronin, Christoph Karp, Guo-Cheng Yuan and Long Cai
2018
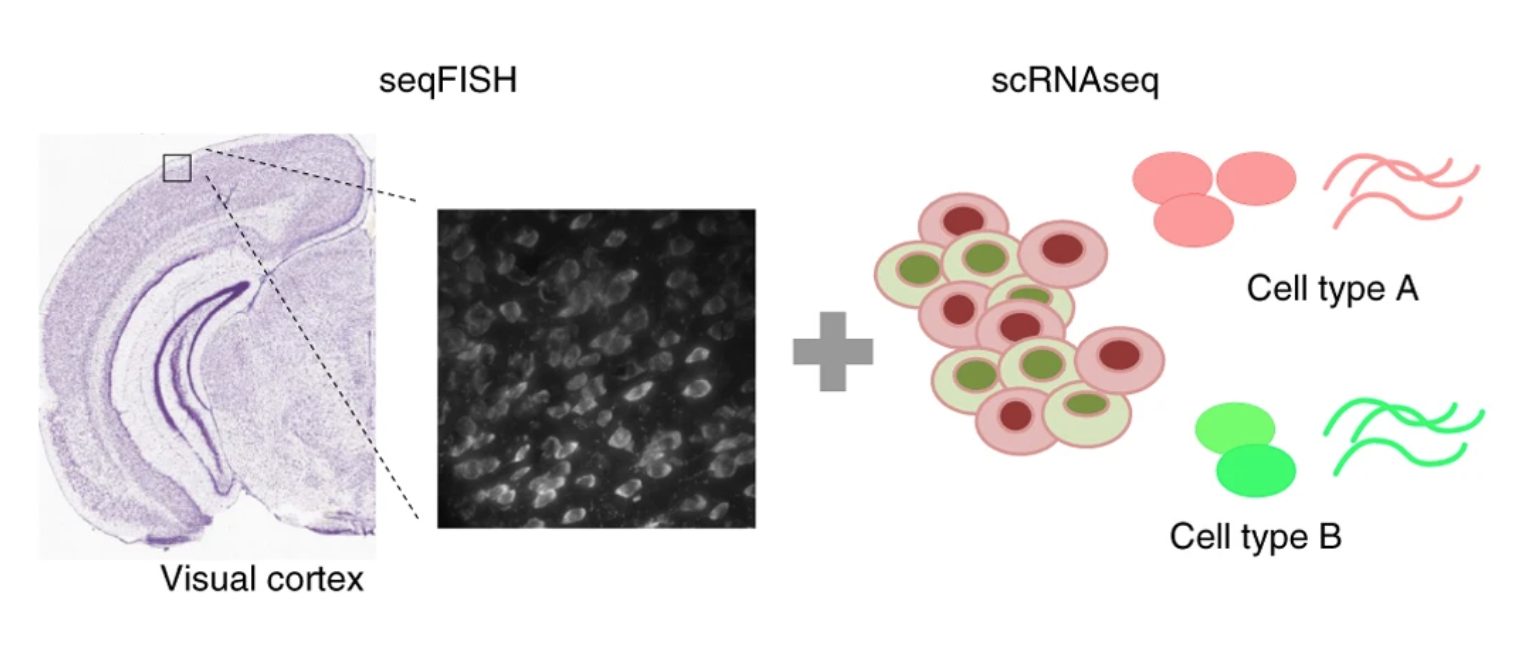
Identification of spatially associated subpopulations by combiningscRNAseq and sequential fluorescence in situ hybridization data
Nature Biotechnology (2018)
Zhu Q, Shah S, Dries R, Cai L*, Yuan GC*

Dynamics and Spatial Genomics of the Nascent Transcriptome by Intron seqFISH
Cell (2018)
Sheel Shah, Yodai Takei, Wen Zhou, Eric Lubeck, Jina Yun, Chee-Huat Linus Eng, Noushin Koulena, Christopher Cronin, Christoph Karp, Eric J. Liaw, Mina Amin and Long Cai
2017
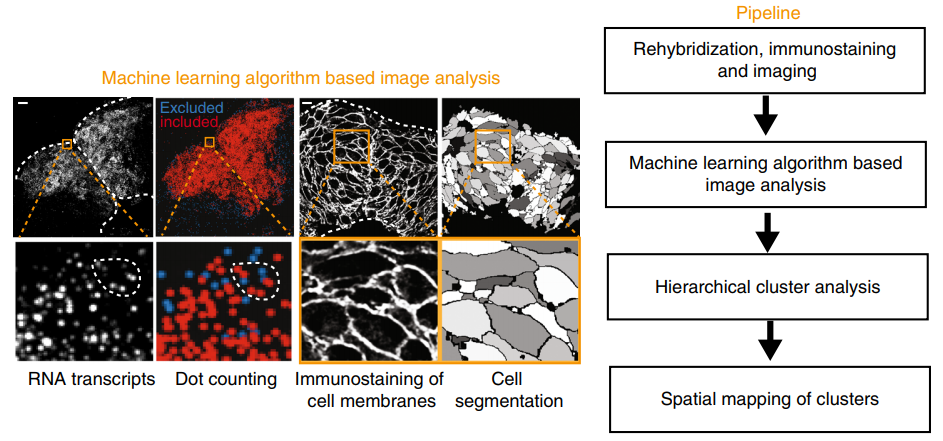
Identification of a neural crest stem cell niche by Spatial Genomic Analysis
Nature Communications (2017)
Antti Lignell, Laura Kerosuo, Sebastian J. Streichan, Long Cai and Marianne E. Bronner

Profiling the transciptome with RNA SPOTs
Nature Methods (2017)
CH. L. Eng, S. Shah, J. Thomassie and L. Cai

seqFISH Accurately Detects Transcripts in Single Cells and Reveals Robust Spatial Organization in the Hippocampus
Neuron (2017)
CH. L. Eng, S. Shah, J. Thomassie and L. Cai

Multiplexed Dynamic Imaging of Genomic Loci by Combined CRISPR Imaging and DNA Sequential FISH
Biophysical Journal (2017)
Y. Takei, S. Shah, S. Harvey, L. S. Qi and L. Cai
2016
Synthetic recording and in situ readout of lineage information in single cells
Nature (2016)
K.L. Frieda+, J.M. Linton+, S. Hormoz+, J. Choi, K.-H.K. Chow, Z.S. Singer, M.W. Budde, M.B. Elowitz* and L. Cai*
In Situ Transcription Profiling of Single Cells Reveals Spatial Organization of Cells in the Mouse Hippocampus
Neuron (2016)
S. Shah+, E. Lubeck+, W. Zhou and L. Cai
Noncommutative Biology: Sequential Regulation of Complex Networks
PLOS Computational Biology (2016)
W. Letsou and L. Cai
Single-molecule RNA detection at depth via hybridization chain reaction and tissue hydrogel embedding and clearing
Development (2016)
S. Shah+, E. Lubeck+, M. Schwarzkopf+, T. He+, A. Greenbaum+, C. H. Sohn, A. Lignell, H. M. T. Choi, V. Gradinaru*, N. A. Pierce* and L. Cai*

Dense transcript profiling in single cells by image correlation decoding
Nature Methods (2016)
A.F. Coskun and L. Cai
2015
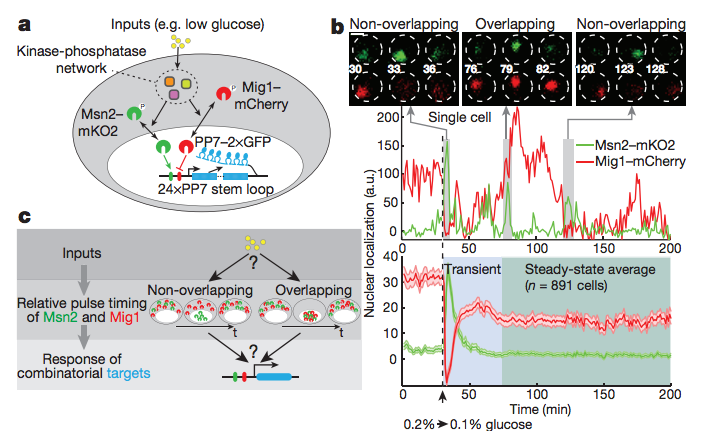
Combinatorial gene regulation by modulation of relative pulse timing
Nature 527, 54-58 (2015)
Y. Lin, C.H. Sohn, C.K. Dalal, L. Cai, M.B. Elowitz
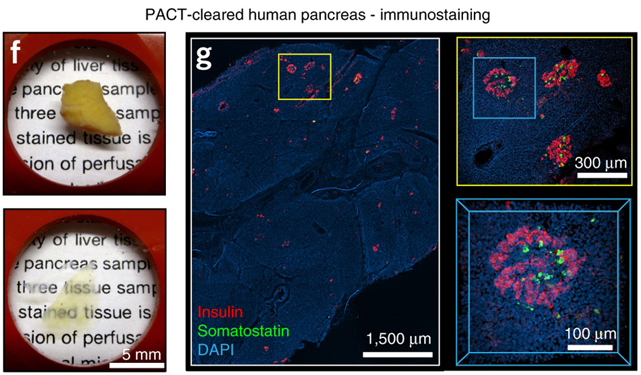
Whole-body tissue stabilization and selective extractions via tissue-hydrogel hybrids for high-resolution intact circuit mapping and phenotyping
Nature Protocols 10, 1860–1896 (2015)
J.B. Treweek,K.Y.Chan,N.C. Flytzanis, B. Yang, B.E. Deverman, A. Greenbaum, A. Lignell, C. Xiao, L.Cai, M.S. Ladinsky,P.J. Bjorkman, C.C. Fowlkes, and V. Gradinaru
2014

Single-cell phenotyping within transparent intact tissue through whole-body clearing
Cell 158, 945 (2014)
B. Yang, J.B. Treweek, R.J. Kulkarni, B.E. Deverman, C. Chen, E. Lubeck,S. Shah, L. Cai, and V. Gradinaru

Dynamic Heterogeneity and DNA Methylation in Embryonic Stem Cells
Molecular Cell 55, 319 (2014)
Z. S. Singer, J. Yong, J. Tischler, J. A. Hackett, A. Altinok, M. A. Surani, L. Cai, and M.B. Elowitz

Single-cell in situ RNA profiling by sequential hybridization
Nature Methods 11, 360 (2014)
E. Lubeck*, A. F. Coskun*, T. Zhiyentayev, M. Ahmad, and L. Cai
2013

Turning single cells into microarrays by super-resolution barcoding
Briefings in Functional Genomics 12, 75 (2013)
L. Cai
*denotes equal contribution
